October 21, 2019
In early September, the version 2.0.0 of rmangal was approved by rOpenSci, four weeks later it made it to CRAN. Following-up on our experience we detail below the reasons why we wrote rmangal, why we submitted our package to rOpenSci and how the peer review improved our package.
 Mangal, a database for ecological networks
Mangal, a database for ecological networks
Ecological networks are defined as a set of species populations (the nodes of the network) connected through ecological interactions (the edges). Interactions are ecological processes in which one species affects another. Although predation is probably the most known and documented interaction, other less noticeable associations are just as essential to ecosystem functioning. For instance, a mammal that unintentionally disperses viable seeds attached to its fur might help plants to thrive. All of these interactions occur simultaneously, shaping ecosystem functioning and making them as complex as they are fascinating.
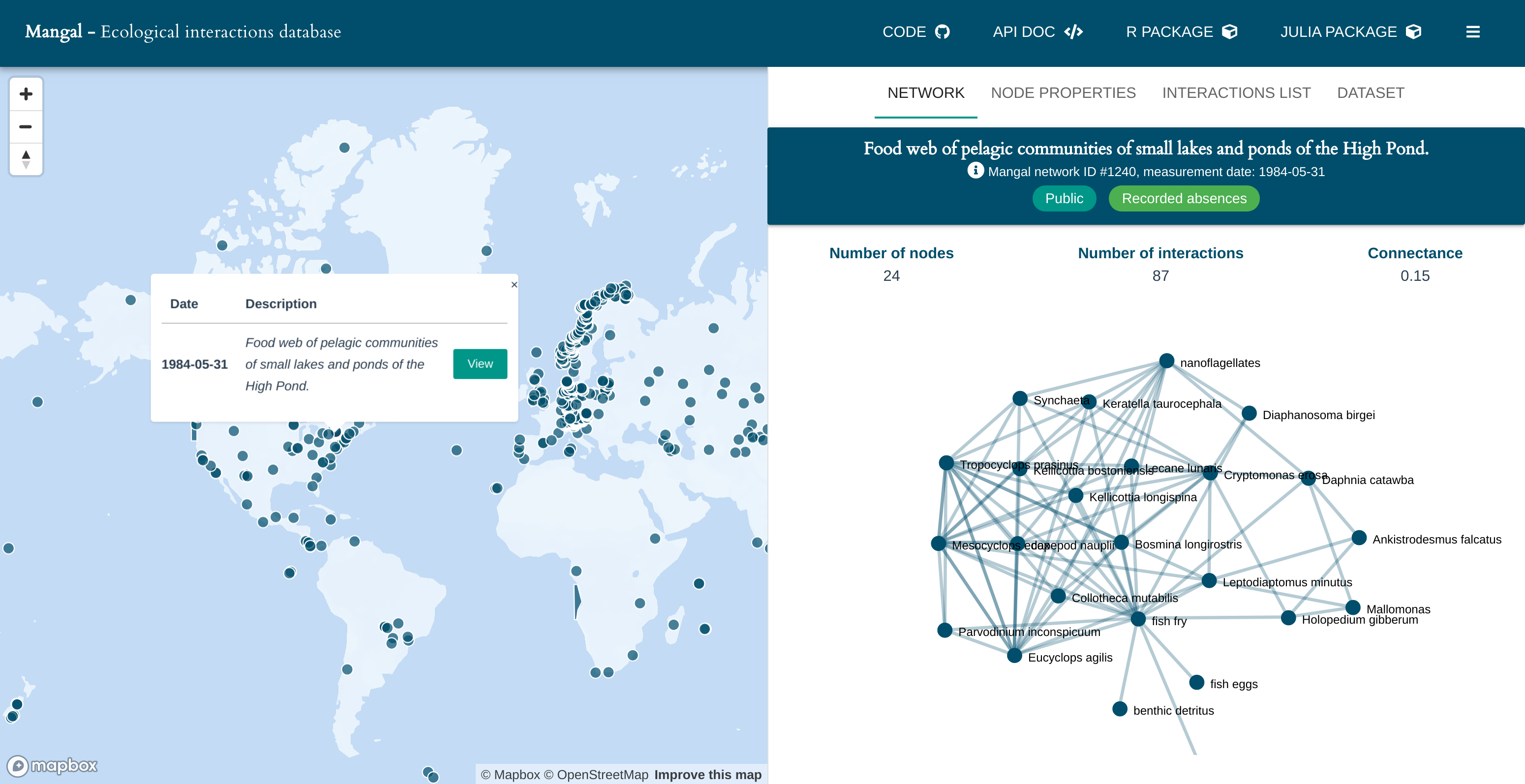
Recording and properly storing these interactions help ecologists to better understand ecosystems. That is why they are currently compiling datasets to explore how species associations vary over environmental gradients and how species lost might affect ecosystem functioning. This fundamental research question should help us understanding how ecological networks will respond to global change. To this end, the Mangal project https://mangal.io/#/ standardizes ecological networks and eases their access. Every dataset contains a collection of networks described in a specific reference (a scientific publication, a book, etc.). For every network included in the database, Mangal includes all the species names and several taxonomic identifiers (gbif, eol, tsn etc.) as well as all interactions and their types. Currently, Mangal includes 172 datasets, which represents over 1300 ecological networks distributed worldwide.
 An R client to make ecological networks easily accessible
An R client to make ecological networks easily accessible
In 2016, the first paper describing the project was published1. In 2018, a substantial effort was made in order to improve the data structure and gather new networks from existing publications. In 2019, the web API was rewritten, a new website launched and hundreds of new interactions were added.
Because of all these modifications, the first version of rmangal was obsolete and a new version needed. It is worth explaining here why the R client is an important component of the Mangal project. Even though Mangal has a documented RESTful API, this web technology is not commonly used by ecologists. On the contrary, providing a R client ensures that the scientific community that documents these interactions in the field can access them, as easily as possible. The same argument holds true for the Julia client that Timothée Poisot wrote because Julia is increasingly popular among theoreticians, that can test ecological theory with such datasets.
We had two main objectives for rmangal 2.0.0. First, the rmangal package had to allow users to search for all entries in the database in a very flexible way. From a technical point this means that we had to write functions to query all the endpoints of the new web API. The second goal was to make the package as user friendly as possible. To do so, we used explicit and consistent names for functions and arguments. We then designed a simple workflow, and documented how to use other field related packages (such as igraph) to visualize and analyze networks (see below). You can find further details in the vignette “get started with rmangal”.
# Loading dependancies
library(rmangal)
library(magrittr)
library(ggraph)
library(tidygraph)
# Retrieving all ecological networks documented in Haven, 1992
havens_1992 <- search_references(doi="10.1126/science.257.5073.1107")
%>% get_collection()
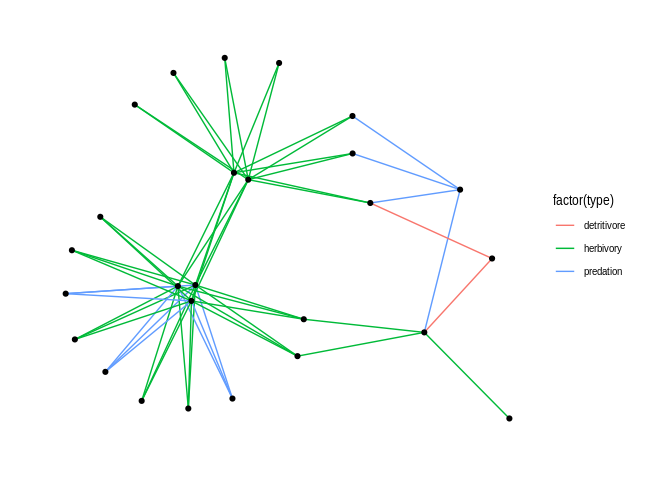
# Coerce and visualize the first network object return by mangal with ggraph
ggraph(as_tbl_graph(havens_1992[[1]])) +
geom_edge_link(aes(colour = factor(type))) +
geom_node_point() +
theme_graph(background = "white")
 A successful peer review process
A successful peer review process
After some hard work behind the screen and once we deemed our two objectives achieved, we decided to submit the rmangal package to rOpenSci for peer review. We did so because we needed feedback, we needed qualified people to critically assess whether our two main objectives were achieved. Given the strong expertise of rOpenSci in software review, and given that our package was in-scope, submitting rmangal to rOpenSci was an obvious choice.
We had very valuable
feedback from Anna
Willoughby and Thomas Lin Pedersen. They carefully assessed
our work and pointed out areas where improvement was required. One good example
of how their review made our package better concerns the dependencies. We
originally listed sf in Imports as we used it to filter networks based
on geographic coordinates. But the reviewers pointed out that this was not an
essential part of the package and that sf has several dependencies. This made us realize that for one extra feature, we were substantially
increasing the number of indirect dependencies. Following the reviewers’
suggestions, we moved sf to Suggests and added a message to warn users
that to use the spatial filtering
feature
requires sf to be installed. Similarly, based on another good comment, we
added a function to convert Mangal networks into tidygraph objects
and we documented how to plot Mangal networks with ggraph (and so we
added those packages in Suggests). Such improvements were very helpful to
properly connect rmangal to the existing R packages. The plethora of R packages
is one of its major strengths, and connecting a package properly to others makes
the entire ecosystem even stronger.
We are now looking for user experience feedback, not only for rmangal (vignette) but also for the web API (documentation) and the mangal.io website. We welcome suggestions and contributions, especially for the documentation by opening new issues on GitHub (mangal-api, mangal-app, rmangal). In the future, we envision that rmangal will integrate functions to format ecological networks for ecologists willing to add their datasets to Mangal. This will likely be the next major release of rmangal.
 Acknowledgments
Acknowledgments
We are thankful to all contributors to rmangal and to all ecologists that have spent countless hours in collecting data. We would like to thank Anna Willoughby and Thomas Lin Pedersen for thorough reviews as well as Noam Ross for handling the review process.
-
Poisot, T. et al. mangal - making ecological network analysis simple. Ecography 39, 384–390 (2016). https://doi.org/10.1111/ecog.00976 ↩︎